|
1/13/2024 0 Comments Genodive input file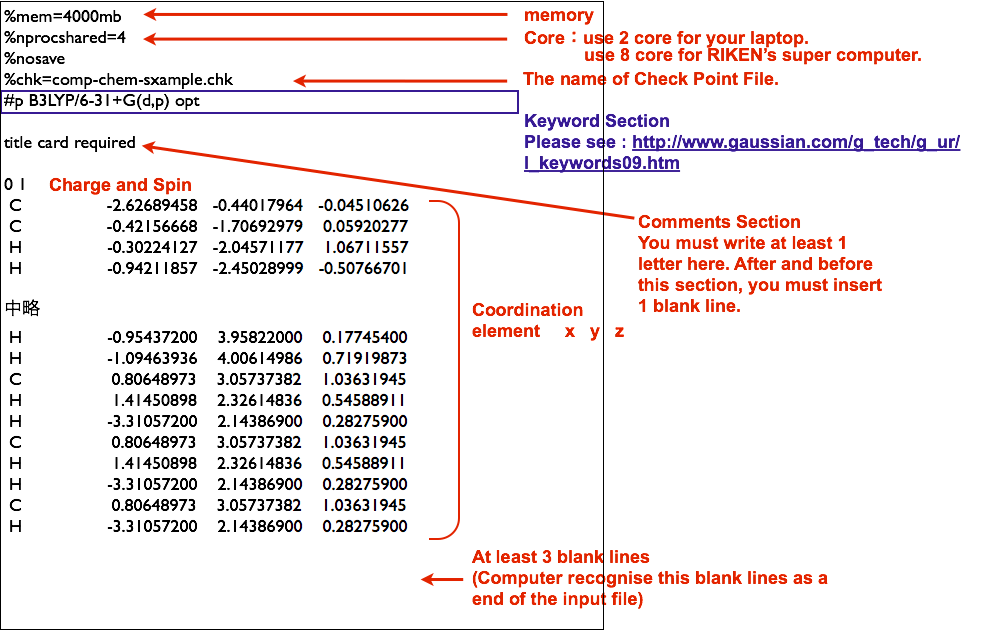
was used as an input file to STRUCTURE HARVESTER (. The field below now shows the file name as well. datasets for analysis using GenoDive 2.0 beta version (Meirmans and. If no file is selected yet, the value is an empty string ( '' ). querySelector ( 'input' ) // Create a new File object const myFile = new File (, 'myFile.txt', A file inputs value attribute contains a string that represents the path to the selected file (s). Get a reference to our file input const fileInput = document. 
We then use DataTransfer to wrap the file in a FileList which we can assign to the files property of our file input element. The script creates a new File object and stores it in myFile We’ll set up a file input element and a short script that sets the file input files property. genodive automatically recognizes which format a text file is in and reads the data, without any need for dialogues. This circumvents tedious recoding of text files from one format to the other, or the use of third-party tools to do such reformatting. The moment we can set the value of a file input the backend seizes to be part of the equation, meaning we can finally progressively enhance the file input element. genodive can seamlessly import and export data in a wide variety of formats. In all these situations there’s no way to upload File objects created in the browser. The backend is so old that literally no one wants to touch it.There’s simply no time or people available to make the required backend changes.We built a web component and don’t know where it’s going to be used. To start simple, lets examine the input file for the monpop data set containing 694 isolates of the plant pathogen Monilinia fructicola.examination of trace files, and by using the Heidelberger-and-Welch diagnostics. By Karine Leydet (5068547), Carsten Grupstra (5068544), Rafel Coma (106536), Marta Ribes (106538) and Michael Hellberg (3309795) Cite. We’re using a NoCode platform or service. INTO FOUR REGIONS IDENTIFIED BY K-MEANS CLUSTERING OF MUS IN GENODIVE.But there are a lot of situations where we don’t control the backend. When we’re in control of the backend we can create an API to handle asynchronous file uploads or convert base64 encoded files back to file ojects. 
We can serve the files as downloads, upload them asynchronously, or convert them to base64 strings and store them in a hidden text field, but we couldn’t store them in a file input element. In both situations we end up with File objects that we can’t leave anywhere but in memory. If you’re interested in why this is useful, read on below, else jump to the solution Why Is Setting The Value Of A File Input Useful?īeing able to set the value of a file input comes in handy when we’ve edited an image in the browser, or when one or more files have been dropped on the page and we want to upload them to a server. From what I learned, sample sizes of n=1 can never be significant.Īlso I calculated Fst values with GenoDive, wich is leading to slightly different Fst values (mean diference 0.0151, sd=0.0699).It’s always been impossible to set the value of a file input element with JavaScript, but this has changed last year and it’s now broadly supported, let’s see how it’s done. read the file into R, and calculate Euclidean distance matrix using 'dist' function (the matrix was called 'd') 2. read.GenoDive takes a text file in the format for the software GenoDive and produces a genambig object. How is that possible? I did check the input file, no problem there. Use FileReader API to read files locally. 
Loop through each file from change event, filter for desired validation. Steps: Add normal file input change event listener. Fst values of "A" compared to other populations shows significant p-values (p<0.05). Below is a round about way of completely avoiding needing to modify the FileList. However I have a population "A" with only one individual. I did calculate Fst values with 3 microsatellite makers for 29 populations using Arlequin 3.5.1.3 (Population pairwise Fst values: Compute F-statistics on haplotype frequencies only).
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
